Illustration by Virginia Greene.
Physiological processes involve dynamic and highly specific cell signaling occurring on a millisecond time scale and largely undetectable in real time through scientific observation. The advent of green fluorescent protein (GFP) and other optical molecules stably expressed under cell-specific transcriptional control in genetically modified organisms has ushered in a new era of in vivo biology allowing for the observation and understanding of complex biological events in real time, in vivo, in mammals.
CHROMusTM will develop mouse lines expressing genetically encoded calcium indicators (GECIs) and optogenetic effectors in lineages relevant to cardiac, vascular, lung and blood diseases. The mouse strains created will be designed to allow for inter crossing resulting in co-expression of sensors with discrete emission wavelengths in interacting lineages (e.g. endothelial and smooth muscle cells), as well as optically compatible effector/detector pairs.
Genetically encoded calcium indicators (GECIs):
GECIs measure calcium, the key signal molecule underlying cell functions such as heart, vessel, and airway contraction, lung secretion, autonomic neurotransmission, and immunocyte function. By using GECIs, researchers are now able to perform in vivo and in situ experiments that were impossible using synthetic reagents such as optical dyes. GECIs allow for studies of cardiovascular, lung, and airway inflammation, cardiac and smooth muscle electrical and chemical signaling, endothelial barrier function and control of vascular tone, organellar signaling, tissue repair, and the functional evaluation of stem cell therapies. Mice will be created in which GCaMP and RCaMP sensors are expressed in multiple cell lineages.
Rhodopsin based Optogenetic Tools:
 Optogenetic actuators have had a stunning impact on the understanding of complex neural circuits, and now consist of a mature and diverse set of optogenetic tools that enable the activation of a broad array of cellular processes and biological events. Combined with cell lineage specific promoters, optogenetic tools enable the activation of discrete cells to initiate biological events. Developments in this area have resulted in a palette of color tuned tools that can initiate cellular processes through the activation of second messengers. Multiple lines of mice will be created expressing channelrhodopsin-2 (ChR2), opto-α1AR and opto-β2AR, all under control of cell specific promoters.
Optogenetic actuators have had a stunning impact on the understanding of complex neural circuits, and now consist of a mature and diverse set of optogenetic tools that enable the activation of a broad array of cellular processes and biological events. Combined with cell lineage specific promoters, optogenetic tools enable the activation of discrete cells to initiate biological events. Developments in this area have resulted in a palette of color tuned tools that can initiate cellular processes through the activation of second messengers. Multiple lines of mice will be created expressing channelrhodopsin-2 (ChR2), opto-α1AR and opto-β2AR, all under control of cell specific promoters.
Combinatorial experiments:
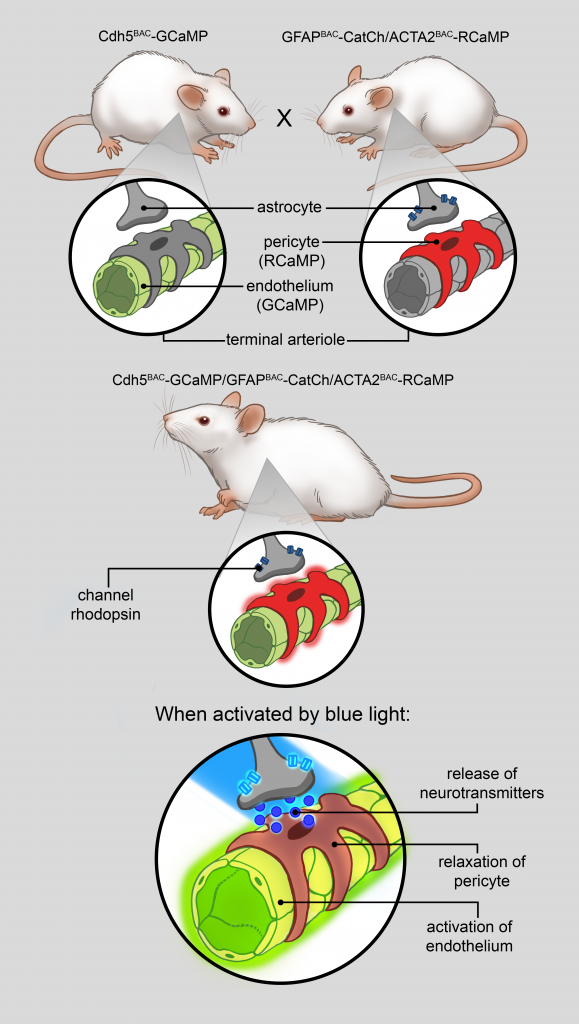
Mouse lines are being developed with the strategic aim of enabling the investigation of cell-cell and subcellular signaling using wavelength-shifted indicator pairs in adjoining lineages or compartments. By crossing two lines together interactions and signaling between two cell lineages can be observed and measured. As an example, a flt1-aOpto1mouse crossed to an acta2-RCaMP1.07 mouse would result in animals in which endothelial cells are triggered by light generating second messenger InsP3/DAG which in turn would activate smooth muscle cells, resulting in an increase in red fluorescence via RCaMP1.07. In a second example, an acta2-RCaMP1.07 mouse crossed with an aMHC-GCaMP8 mouse would allow investigators to examine endothelial cell/smooth muscle cell signaling in vivo by looking at the linkage between the calcium responses in each tissue, which are coded by color. Other pairs of mice can be combined to study arrhythmias by triggering ectopic cardiac activation of heart cells with light pulses and looking at the effects on heart activation and nodal signaling. Other crosses would enable examination of the link between heart cell function and blood flow.
Transgenic constructs:
Mice will be created with BAC constructs or well characterized minimal promoters (e.g. αMHC) that confer tight tissue specificity, and lines screened for lineage restriction, high expression, and optical function. Many promoters have been extensively studied to determine the minimal DNA sequence needed for basic expression. Unfortunately fidelity and maximum expression levels are not always observed with minimal promoters. Using BAC recombineering technology we will be able to ensure tight tissue specificity. For details on the construction of transgenes, click here.
